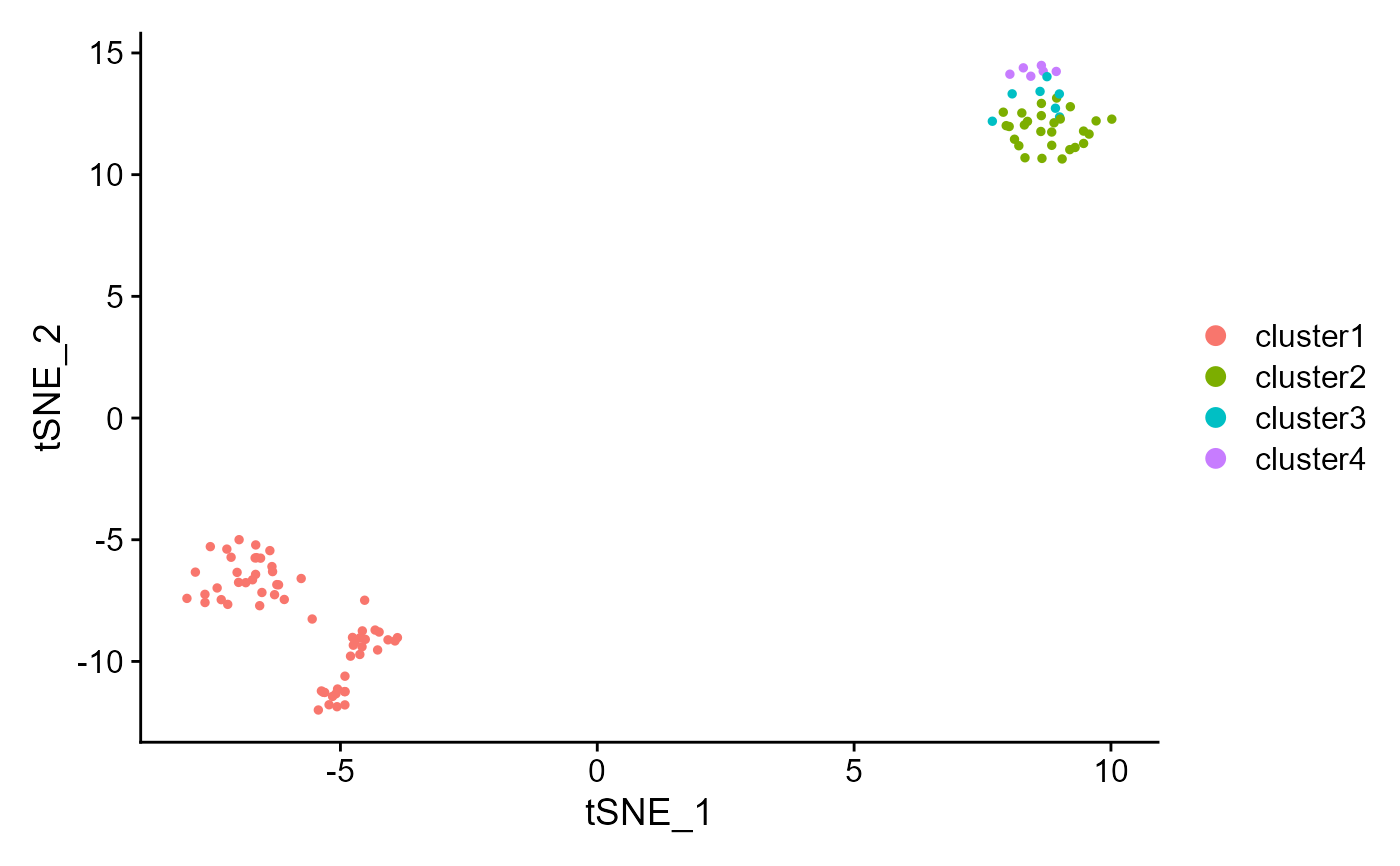
tNSE or UMAP plot visualization
drscPlot.RdIntuitive way of visualizing how cell types changes across the embeddings obatined by DR-SC.
drscPlot(seu, dims=1:5, visu.method='tSNE',...)Arguments
Details
Nothing
Value
return a ggplot2 object.
References
None
Note
nothing
See also
None
Examples
## we generate the spatial transcriptomics data with lattice neighborhood, i.e. ST platform.
data(seu)
library(Seurat)
seu <- NormalizeData(seu)
#> Normalizing layer: counts
# choose spatially variable features
seu <- FindSVGs(seu)
#> Find the spatially variables genes by SPARK-X...
#> ## ===== SPARK-X INPUT INFORMATION ====
#> ## number of total samples: 100
#> ## number of total genes: 50
#> ## Running with single core, may take some time
#> ## Testing With Projection Kernel
#> ## Testing With Gaussian Kernel 1
#> ## Testing With Gaussian Kernel 2
#> ## Testing With Gaussian Kernel 3
#> ## Testing With Gaussian Kernel 4
#> ## Testing With Gaussian Kernel 5
#> ## Testing With Cosine Kernel 1
#> ## Testing With Cosine Kernel 2
#> ## Testing With Cosine Kernel 3
#> ## Testing With Cosine Kernel 4
#> ## Testing With Cosine Kernel 5
# use SVGs to fit DR.SC model
# maxIter = 2 is only used for illustration, and user can use default.
seu1 <- DR.SC(seu, K=4,platform = 'ST', maxIter = 2,verbose=FALSE)
#> Neighbors were identified for 100 out of 100 spots.
#> Fit DR-SC model...
#> Using accurate PCA to obtain initial values
#> Finish DR-SC model fitting
drscPlot(seu1)